neural posterior estimation for population genetics
With Jiseon Min, Nathaniel Pope, and Andrew Kern we worked on a workflow that enables simulation based machine learning inference for population genetics. popgen-npe infers population genetic parameters using neural posterior estimation, enabling likelihood-free analysis of complex evolutionary models in a Bayesian framework.
If you want to try it go to github.
identifying sampling bias in bacterial genome databases

When analyzing the diversity of bacterial populations it is important to consider a potential sampling bias that can distort your pangenome analysis.
Our tool PhyloThin is a coalescent-theory-based approach to identify prokaryotic genomes that can be considered as oversampled.
We are currently writing the manuscript for this tool. If you need preliminary access to PhyloThin, check out our github repository.
ancestral reconstruction of phage infection histories
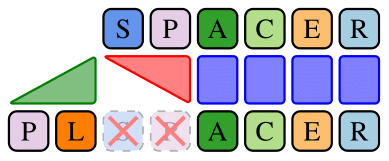
The immunity memory of the prokaryotic defense system CRISPR-Cas is remarkable, as it encodes a chronological record of past infection attempts in an inheritable spacer array. SpacerPlacer is a tool designed to reconstruct the ancestral states and events that formed this spacer array. If you want to reconstruct the spacer arrays of your favorite species, check out our GitHub repository. We reconstructed the deletion events of spacers in the CRISPRCasdb database, revealing some interesting properties of immunity loss in CRISPR-Cas systems. Have a look at our paper to find out more.
phylogeny-aware detection of gene associations and phage-pangenome interactions

Goldfinder identifies co-occurring and mutually exclusive genes in bacterial pangenomes while accounting for shared evolutionary history. It can also be used to identify interactions between prokaryotic genes and phages.
effective large scale coalescent simulations

msprime uses succinct tree sequences to speed up the simulation of ancestral relationships. We are part of the fantastic scientific open-source community that develops novel features into msprime maintained by Jerome Kelleher. We have implemented the gene conversion mechanism into msprime and currently incorporate additional features that are particularly relevant for microbial evolution.
Have a look at the msprime documentation here.
supervised feature selection for ancestry informative markers
With Peter Pfaffelhuber, Franziska Grundner-Culemann and Veronika Lipphardt we created AIMsetfinder.
AIMsetfinder is a supervised feature selection approach to identify sets of Ancestry Informative Markers (AIMs), that minimize the logloss error of a naive Bayes classifier.
Have a look at what we have done at github
standardizing population genetic simulations
we are a member of the PopSIm Consortium, aiming to standardize population genetic simulations for frequently studied model organisms and species, including humans.
As a result, the software stdpopsim allows to a) easily re-simulate published population models for many species and b) compare the results of new inference methods against a standardized benchmark.
Have a look at the stdpopsim documentation here.
Seite 1 von 2
